Mvp-objects¶
One of skbold’s main features is the data-structure Mvp (an abbreviation
of MultiVoxel Pattern). This custom object allows for an efficient way
to store and access data and metadata necessary for multivoxel analyses of fMRI data.
A nice feature of this Mvp objects is that they can easily load data
(i.e., sets of nifti-files) from disk and automatically organize it in
a format that is used in ML-analyses (i.e., a sample-by-feature matrix).
So, at the core, an Mvp-object is simply a collection of data - a 2D array
of samples by features - and fMRI-specific metadata necessary to perform
customized preprocessing and feature engineering. However, machine learning
analyses, or more generally any type of multivoxel-type analysis (i.e. MVPA),
can be done in two basic ways, which provide the basis of the two ‘flavors’
of Mvp-objects: MvpWithin and MvpBetween, as explained in more
detail below.
MvpWithin¶
One way is to perform analyses within subjects. This means that a model is fit on each subjects’ data separately. Data, in this context, often refers to single-trial data, in which each trial comprises a sample in our data-matrix and the values per voxel constitute our features. This type of analysis is alternatively called single-trial decoding, and is often performed as an alternative to (whole-brain) univariate analysis.
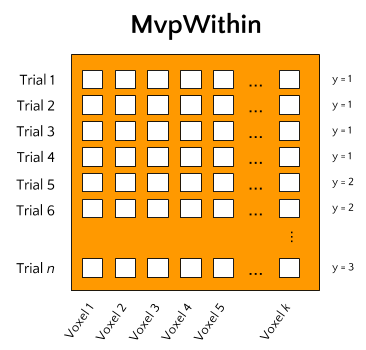
Ultimately, this type of analysis aims to predict some kind of attribute of the
trials (for example condition/class membership in classification analyses or some
continuous feature in regression analyses), which skbold calls y, based
on a model trained on the samples-by-features matrix, which skbold calls X.
After obtaining model performance scores (such as accuracy, F1-score, or R-squared)
for each subject, a group-level random effects (RFX) analysis can be done on
these scores. Skbold does not offer any functionality in terms of group-level
analyses; we advise researchers to look into the
prevalance inference
method of Allefeld and colleagues.
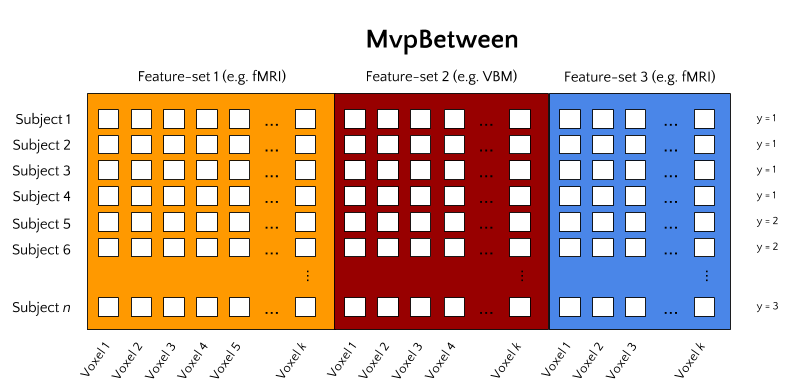
MvpBetween¶
With the increase in large-sample neuroimaging datasets, another type of MVPA starts to become feasible, which we’ll call between subject analyses. In this type of analysis, single subjects constitute the data’s samples and a corresponding single multivoxel pattern constitutes the data’s features. The type of multivoxel pattern, or ‘feature-set’, can be any set of voxel values. For example, features from a single first-level contrast (note: this should be a condition average contrast, as opposed to single-trial contrasts in MvpWithin!) can be used. But voxel patterns from VBM, TBSS (DTI), and dual-regression maps can equally well be used. Crucially, this package allows for the possibility to stack feature-sets such that models can be fit on features from multiple data-types simultaneously.